# Static correlation heatmap - one view only
data(mtcars)
cor_mtcars <- cor(mtcars)
# Choose one color scheme, one ordering, one threshold
corrplot(cor_mtcars, method = "circle", order = "hclust",
tl.col = "black")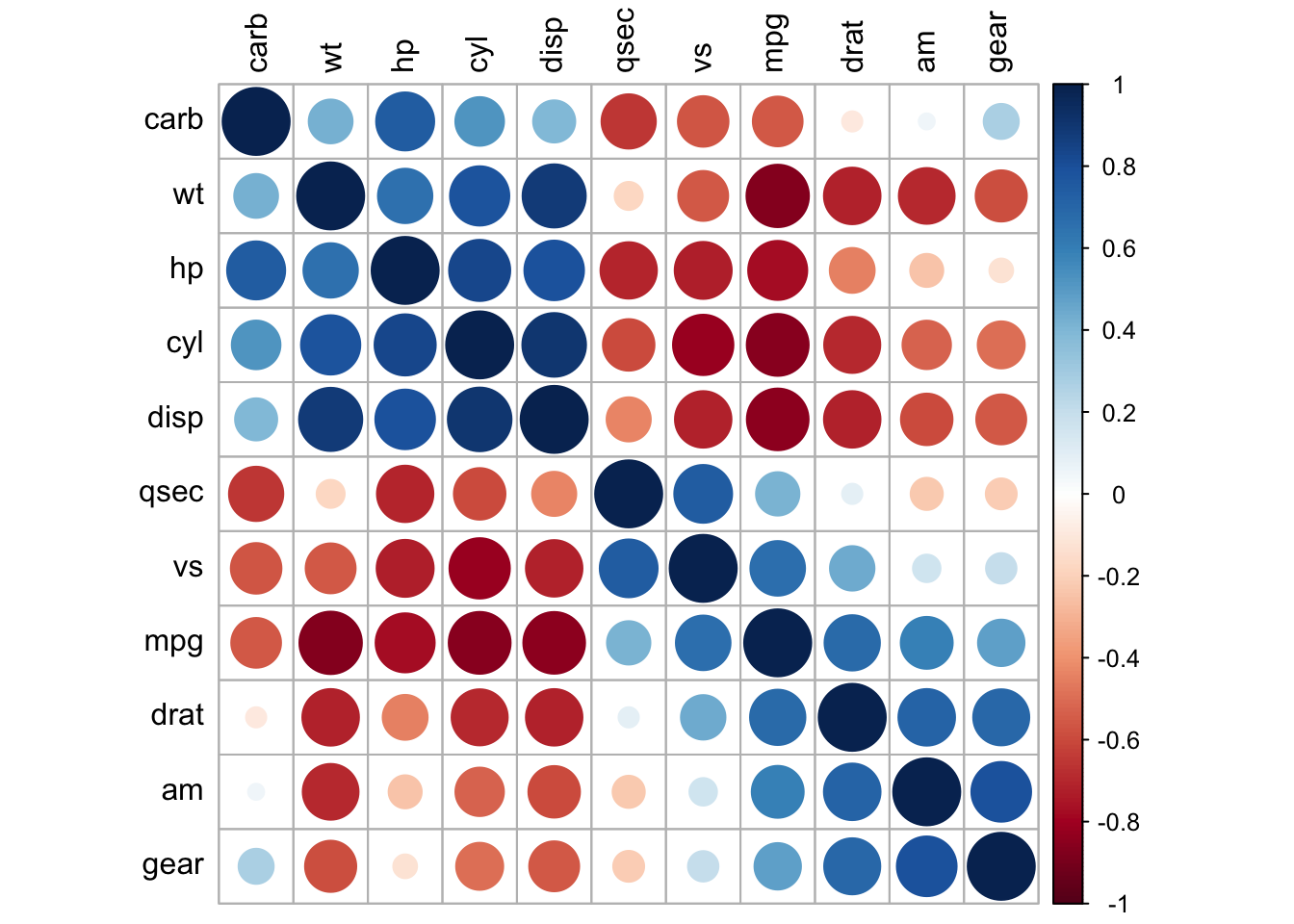
In previous chapters, we computed association matrices, created correlation heatmaps, and drew network graphs. These static visualizations are publication-ready and reproducible, but they can be inherently limited: each figure represents one choice of parameters, one threshold, one layout, one subset of data.
Consider a researcher working with network data:
Static analysis answers one question at a time. Each new question typically requires code modification, recomputation, and a new figure.
Interactive analysis can empower exploration. Users can adjust parameters, filter data, and compare perspectives without waiting for recomputation, enabling discovery of patterns that static workflows often miss.
This chapter introduces Shiny, an R web framework that can transform exploratory analysis from a batch workflow into an interactive conversation with data. Rather than asking “what does this plot show?”, interactive tools let practitioners ask “what if I change this parameter?” or “how does this look in this subgroup?” and receive near-instant visual feedback.
A single figure is frozen in time and parameter space. Consider a correlation matrix visualization:
# Static correlation heatmap - one view only
data(mtcars)
cor_mtcars <- cor(mtcars)
# Choose one color scheme, one ordering, one threshold
corrplot(cor_mtcars, method = "circle", order = "hclust",
tl.col = "black")
This figure effectively communicates the correlation structure, but:
Practitioners often resort to creating multiple figures to show different perspectives, one for each threshold, subgroup, or parameter choice. This multiplies the number of static outputs and makes comparison harder, not easier.
Interactive tools address these limitations by enabling:
These capabilities can transform exploration from passive viewing to active interrogation.
Interactive exploration is most valuable when:
Conversely, we still need static visualizations for publication and presentation, where reproducibility and editorial control matter more than exploration.
Shiny separates an application into two components:
A minimal Shiny app has this structure:
library(shiny)
# Define UI
ui <- fluidPage(
titlePanel("My First Shiny App"),
sidebarLayout(
sidebarPanel(
sliderInput("threshold", "Correlation Threshold:",
min = 0, max = 1, value = 0.5, step = 0.05)
),
mainPanel(
plotOutput("correlation_plot")
)
)
)
# Define server logic
server <- function(input, output) {
output$correlation_plot <- renderPlot({
# This code runs whenever input$threshold changes
cor_mtcars <- cor(mtcars)
# Create network based on current threshold
adj <- (abs(cor_mtcars) > input$threshold) * 1
diag(adj) <- 0
# Create visualization...
})
}
# Run the app
shinyApp(ui, server)The key concept is that when a user adjusts the slider, the value in input$threshold updates, triggering a recomputation of output$correlation_plot.
Reactivity is Shiny’s mechanism for connecting inputs to outputs. A reactive expression recomputes only when its dependencies change:
server <- function(input, output) {
# Reactive expression: computes only when threshold changes
net_data <- reactive({
cor_mtcars <- cor(mtcars)
adj <- (abs(cor_mtcars) > input$threshold) * 1
diag(adj) <- 0
graph_from_adjacency_matrix(adj, mode = "undirected", weighted = TRUE)
})
# Outputs can depend on reactive expressions
output$net_viz <- renderPlot({
net <- net_data()
plot(net)
})
output$summary_stats <- renderTable({
net <- net_data()
tibble(
Metric = c("Nodes", "Edges", "Density"),
Value = c(vcount(net), ecount(net), edge_density(net))
)
})
}Notice: net_data() is defined once but used in multiple outputs. Both outputs automatically update together when input$threshold changes. This dependency tracking is automatic in Shiny (you need not explicitly code update sequences).
Shiny provides many input types:
# Slider for continuous values
sliderInput("threshold", "Threshold:", min = 0, max = 1, value = 0.5)
# Select dropdown for choices
selectInput("variable", "Variable:", choices = c("x1", "x2", "x3"))
# Radio buttons for mutually exclusive options
radioButtons("method", "Method:",
choices = c("Pearson", "Spearman", "Kendall"))
# Checkbox for binary options
checkboxInput("show_labels", "Show node labels", value = TRUE)
# Multiple selection
selectInput("subgroups", "Filter by group:",
choices = unique(data$group),
multiple = TRUE)
# Date range picker
dateRangeInput("date_range", "Select date range:",
start = "2020-01-01", end = "2023-12-31")Similarly, outputs can render various visualizations and tables:
# Plot output
plotOutput("myplot")
# Table output
tableOutput("mytable")
# or for prettier tables:
dataTableOutput("interactive_table")
# Text output
textOutput("summary_text")
# HTML output (for custom formatting)
htmlOutput("custom_html")
# UI output (dynamically generated controls)
uiOutput("dynamic_controls")Let us build a practical interactive tool that explores how network structure changes as the correlation threshold varies:
library(shiny)
library(igraph)
library(tidygraph)
library(ggraph)
library(ggplot2)
library(dplyr)
ui <- fluidPage(
titlePanel("Correlation Network Threshold Explorer"),
sidebarLayout(
sidebarPanel(
sliderInput("threshold",
"Correlation Threshold:",
min = 0, max = 1, value = 0.5, step = 0.05),
helpText("Adjust to see how network structure changes."),
br(),
downloadButton("downloadPlot", "Download Plot")
),
mainPanel(
tabsetPanel(
tabPanel("Network Visualization",
plotOutput("network_plot", height = "600px")),
tabPanel("Network Statistics",
tableOutput("network_stats")),
tabPanel("Correlation Distribution",
plotOutput("cor_distribution"))
)
)
)
)
server <- function(input, output, session) {
# Reactive network data
net_reactive <- reactive({
cor_mtcars <- cor(mtcars)
adj <- (abs(cor_mtcars) > input$threshold) * 1
diag(adj) <- 0
graph_from_adjacency_matrix(adj, mode = "undirected", weighted = TRUE)
})
# Network plot
output$network_plot <- renderPlot({
net <- net_reactive()
net_tidy <- as_tbl_graph(net)
ggraph(net_tidy, layout = 'fr') +
geom_edge_link(edge_colour = "gray60", alpha = 0.5) +
geom_node_point(size = 5, colour = "steelblue") +
geom_node_text(aes(label = name), repel = TRUE, size = 3) +
labs(title = paste("Network at Threshold =", input$threshold)) +
theme_void()
})
# Network statistics table
output$network_stats <- renderTable({
net <- net_reactive()
tibble(
Metric = c("Nodes", "Edges", "Density", "Average Degree", "Diameter"),
Value = c(
vcount(net),
ecount(net),
round(edge_density(net), 3),
round(mean(degree(net)), 3),
ifelse(ecount(net) > 0, diameter(net), NA)
)
)
}, striped = TRUE, bordered = TRUE)
# Distribution of all correlations
output$cor_distribution <- renderPlot({
cor_mtcars <- cor(mtcars)
cor_values <- cor_mtcars[upper.tri(cor_mtcars)]
ggplot(tibble(correlation = cor_values), aes(x = correlation)) +
geom_histogram(bins = 20, fill = "steelblue", alpha = 0.7) +
geom_vline(xintercept = input$threshold, colour = "red", linetype = "dashed", size = 1) +
geom_vline(xintercept = -input$threshold, colour = "red", linetype = "dashed", size = 1) +
labs(title = "Distribution of All Correlations",
subtitle = "Red lines show current threshold",
x = "Correlation", y = "Frequency") +
theme_minimal()
})
# Download handler
output$downloadPlot <- downloadHandler(
filename = function() {
paste0("network_threshold_", input$threshold, ".png")
},
content = function(file) {
png(file, width = 800, height = 600)
net <- net_reactive()
net_tidy <- as_tbl_graph(net)
plot(ggraph(net_tidy, layout = 'fr') +
geom_edge_link(edge_colour = "gray60", alpha = 0.5) +
geom_node_point(size = 5, colour = "steelblue") +
geom_node_text(aes(label = name), repel = TRUE) +
theme_void())
dev.off()
}
)
}
shinyApp(ui, server)Interactive tools excel at comparing patterns across subgroups. Here is an application that compares correlation networks across iris species:
ui <- fluidPage(
titlePanel("Iris Network by Species"),
sidebarLayout(
sidebarPanel(
selectInput("species", "Select Species:",
choices = unique(iris$Species)),
sliderInput("threshold",
"Correlation Threshold:",
min = 0, max = 1, value = 0.5, step = 0.05),
radioButtons("layout", "Graph Layout:",
choices = c("Force-Directed" = "fr",
"Circle" = "circle"))
),
mainPanel(
plotOutput("species_network", height = "600px"),
br(),
tableOutput("centrality_table")
)
)
)
server <- function(input, output, session) {
# Filter data by species
species_data <- reactive({
iris |>
filter(Species == input$species) |>
select(-Species)
})
# Compute network for selected species
species_net <- reactive({
data <- species_data()
cor_mat <- cor(data)
adj <- (abs(cor_mat) > input$threshold) * 1
diag(adj) <- 0
graph_from_adjacency_matrix(adj, mode = "undirected", weighted = TRUE)
})
# Plot network with user-selected layout
output$species_network <- renderPlot({
net <- species_net()
net_tidy <- as_tbl_graph(net)
layout_func <- if (input$layout == "fr") {
"fr"
} else {
"circle"
}
ggraph(net_tidy, layout = layout_func) +
geom_edge_link(edge_colour = "gray60", alpha = 0.6) +
geom_node_point(size = 6, colour = "coral") +
geom_node_text(aes(label = name), repel = TRUE, size = 4) +
labs(title = paste("Network for", input$species)) +
theme_void()
})
# Node centrality measures
output$centrality_table <- renderTable({
net <- species_net()
tibble(
Variable = names(species_data()),
Degree = degree(net),
Betweenness = round(betweenness(net), 2),
Closeness = round(closeness(net), 3)
) |>
arrange(desc(Degree))
}, striped = TRUE)
}
shinyApp(ui, server)As apps grow complex, they can become slow if computations repeat unnecessarily. The solution is strategic use of reactive expressions:
# Bad: Correlations computed twice
server <- function(input, output) {
output$heatmap <- renderPlot({
cor_data <- cor(mtcars) # Computed here
# plot heatmap
})
output$network <- renderPlot({
cor_data <- cor(mtcars) # Computed again!
# plot network
})
}
# Good: Correlations computed once, reused
server <- function(input, output) {
cor_reactive <- reactive({
cor(mtcars)
})
output$heatmap <- renderPlot({
cor_data <- cor_reactive() # Uses cached result
# plot heatmap
})
output$network <- renderPlot({
cor_data <- cor_reactive() # Reuses same result
# plot network
})
}Some inputs change very rapidly (e.g., a slider dragged quickly). Without debouncing, Shiny recomputes on every drag. Solutions include:
# Debounce: only recompute 0.5 seconds after slider stops moving
threshold_debounced <- debounce(reactive(input$threshold), 500)
server <- function(input, output) {
output$plot <- renderPlot({
# Uses debounced threshold; only recomputes after user stops dragging
threshold <- threshold_debounced()
# ... plot code ...
})
}
# Alternatively, use an "Apply" button to batch updates
server <- function(input, output) {
apply_button <- eventReactive(input$apply_button, {
list(threshold = input$threshold,
species = input$species)
})
output$plot <- renderPlot({
params <- apply_button() # Only recomputes when button is clicked
# ... plot code ...
})
}Apps often need dynamic UI: show certain controls only when relevant. Shiny provides conditionalPanel():
ui <- fluidPage(
sidebarPanel(
selectInput("plot_type", "Plot Type:",
choices = c("Network", "Heatmap", "Scatter")),
# Show network options only when "Network" is selected
conditionalPanel(
condition = "input.plot_type == 'Network'",
sliderInput("threshold", "Threshold:", min = 0, max = 1, value = 0.5),
radioButtons("layout", "Layout:",
choices = c("fr", "circle", "spring"))
),
# Show heatmap options only when "Heatmap" is selected
conditionalPanel(
condition = "input.plot_type == 'Heatmap'",
radioButtons("heatmap_method", "Distance Method:",
choices = c("euclidean", "correlation"))
)
),
mainPanel(
plotOutput("main_plot")
)
)Users can be overwhelmed by too many options at once. It helps to group controls logically and use tabs or collapsible panels:
ui <- fluidPage(
tabsetPanel(
tabPanel("Data Overview",
# Simple view: just load and show data
fileInput("data_file", "Choose CSV"),
tableOutput("data_preview")),
tabPanel("Exploration",
# Intermediate view: threshold, subset options
sliderInput("threshold", "Threshold:", 0, 1, 0.5)),
tabPanel("Advanced",
# Advanced view: algorithms, layouts, statistics
radioButtons("algorithm", "Algorithm:", ...),
radioButtons("layout", "Layout:", ...))
)
)Users often benefit from immediate visual feedback. Try to avoid long computations. Instead:
withProgress() to show a progress bar for longer operationsToo many controls can confuse users. Consider prioritizing:
When comparing across subgroups or time periods, arrange views side-by-side or with linked highlighting:
ui <- fluidPage(
fluidRow(
column(6, h3("Species A"), plotOutput("net_a")),
column(6, h3("Species B"), plotOutput("net_b"))
)
)While powerful, interactive exploration has drawbacks:
Best practice: Use interactive tools for exploration, then document findings in static reports.
The chapters so far have built the foundation for association analysis:
Now we integrate all these components into a cohesive, interactive tool. The next chapter introduces AssociationExplorer, a Shiny application designed specifically for exploratory association analysis. AssociationExplorer combines:
AssociationExplorer demonstrates how the principles and techniques from this chapter, combined with sound statistical practice and user-centered design, enable practitioners to explore high-dimensional association structures interactively, discover patterns that static analysis would miss, and communicate findings effectively to diverse audiences.
With Shiny principles established, we now turn to a complete, production-ready application: AssociationExplorer. The following chapter showcases how these principles integrate into a comprehensive tool for exploring, understanding, and communicating association structures in complex tabular data.