# Create dataset with various nonlinear relationships
set.seed(42)
n <- 200
nonlin_data <- tibble(
Linear = seq(0, 10, length.out = n),
Quadratic = seq(0, 10, length.out = n),
Exponential = seq(0, 10, length.out = n),
Sine = seq(0, 4 * pi, length.out = n),
x_lin = rnorm(n, 0, 1),
x_quad = rnorm(n, 0, 1),
x_exp = rnorm(n, 0, 1),
x_sin = rnorm(n, 0, 1)
) |>
mutate(
y_lin = Linear + x_lin,
y_quad = 10 - Quadratic^2 / 8 + x_quad,
y_exp = exp(Exponential / 3) + x_exp * 2,
y_sin = sin(Sine) + x_sin
)
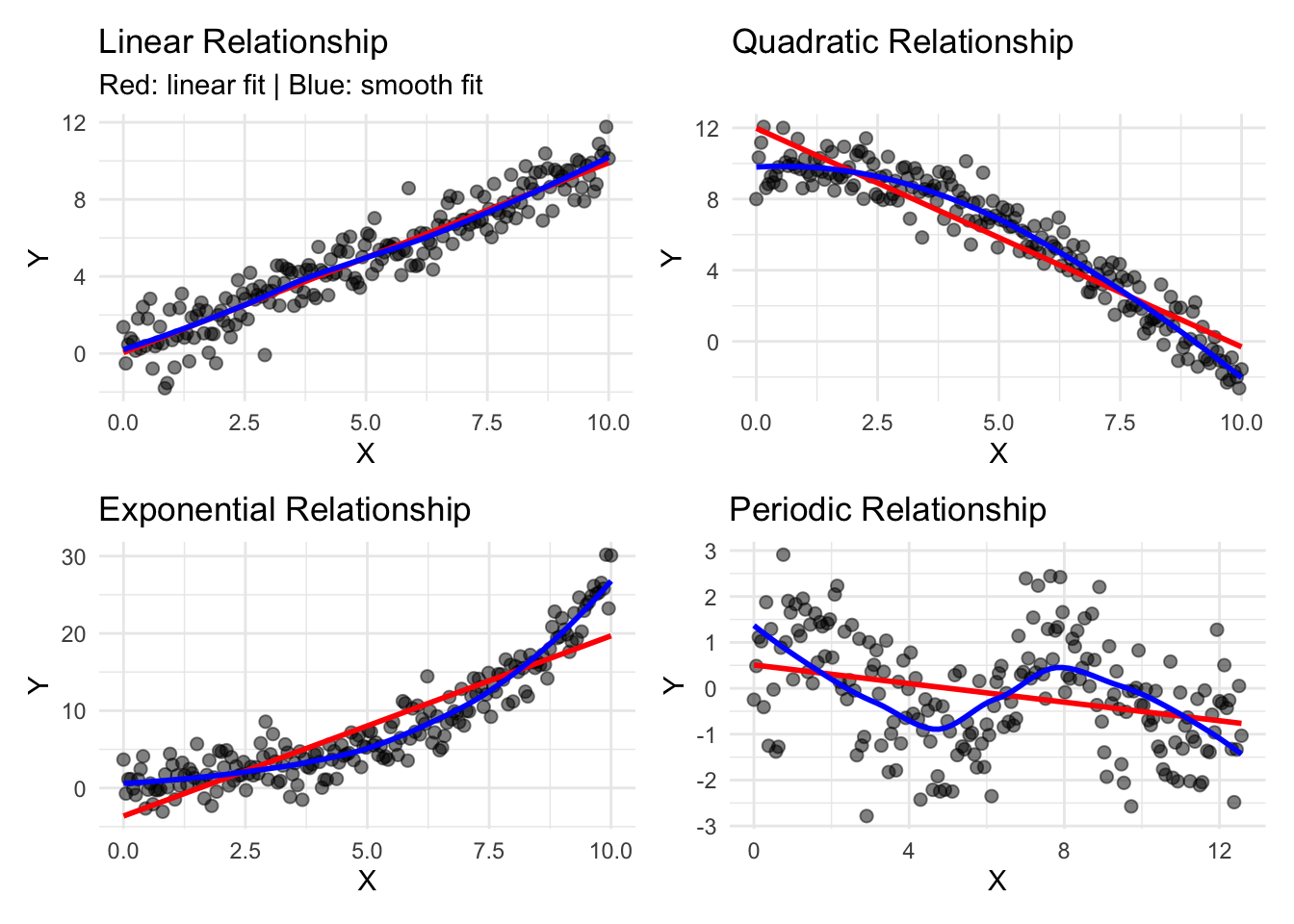
# Visualize multiple relationship types
p1 <- nonlin_data |>
ggplot(aes(x = Linear, y = y_lin)) +
geom_point(alpha = 0.5, size = 2) +
geom_smooth(method = "lm", se = FALSE, color = "red", linewidth = 1) +
geom_smooth(method = "loess", se = FALSE, color = "blue", linewidth = 1) +
labs(
title = "Linear Relationship",
x = "X", y = "Y",
subtitle = "Red: linear fit | Blue: smooth fit"
) +
theme_minimal()
p2 <- nonlin_data |>
ggplot(aes(x = Quadratic, y = y_quad)) +
geom_point(alpha = 0.5, size = 2) +
geom_smooth(method = "lm", se = FALSE, color = "red", linewidth = 1) +
geom_smooth(method = "loess", se = FALSE, color = "blue", linewidth = 1) +
labs(
title = "Quadratic Relationship",
x = "X", y = "Y"
) +
theme_minimal()
p3 <- nonlin_data |>
ggplot(aes(x = Exponential, y = y_exp)) +
geom_point(alpha = 0.5, size = 2) +
geom_smooth(method = "lm", se = FALSE, color = "red", linewidth = 1) +
geom_smooth(method = "loess", se = FALSE, color = "blue", linewidth = 1) +
labs(
title = "Exponential Relationship",
x = "X", y = "Y"
) +
theme_minimal()
p4 <- nonlin_data |>
ggplot(aes(x = Sine, y = y_sin)) +
geom_point(alpha = 0.5, size = 2) +
geom_smooth(method = "lm", se = FALSE, color = "red", linewidth = 1) +
geom_smooth(method = "loess", se = FALSE, color = "blue", linewidth = 1) +
labs(
title = "Periodic Relationship",
x = "X", y = "Y"
) +
theme_minimal()
(p1 + p2) / (p3 + p4)